Eclipse Triquetrum - An open platform for scientific workflows

Source: Wikipedia
What is Eclipse Triquetrum?
Eclipse Triquetrum delivers an open platform for managing and executing scientific workflows. The goal of Triquetrum is to support a wide range of use cases, ranging from automated processes based on predefined models, to replaying ad-hoc research workflows recorded from a user's actions in a scientific workbench UI. It will allow the user to define and execute models from personal pipelines with a few steps to massive models with thousands of elements.
Besides delivering a generic workflow environment, Triquetrum will also deliver extensions with a focus on scientific software. There is no a-priori limitation on target scientific domains, but the currently interested organizations are big research institutions in materials research (synchrotrons), physics and engineering.
Compared to other platforms for scientific workflows, Triquetrum differentiates itself by:
- combining the mature actor-oriented Ptolemy II framework [1] with Eclipse and OSGi technology
- being designed explicitly for integration in larger scientific software systems
The combination of Eclipse/OSGi with Ptolemy II delivers a solid platform for a wide range of workflow applications, and in particular for scientific workflows. The modularity and dynamism offered by OSGi, the rich set of frameworks and technologies offered through the Eclipse Foundation, and the community of the Eclipse Science Working Group together result in a very powerful ecosystem for projects like Triquetrum.
The project is structured on three lines of work:
- Workflow-related features:
- an execution runtime that must be easy to integrate in different environments, ranging from a personal RCP workbench to large-scale distributed systems.
- a graphical model editor delivered as plugins for an Eclipse RCP application.
- APIs and OSGi service implementations for task-based processing. This is a Triquetrum subsystem that is designed to be reusable independently of the workflows as well. It can easily be integrated in other process-oriented applications to decompose processes in sequences of traceable steps, including support for concurrent and asynchronous work.
- Supporting APIs and tools, e.g. integration adapters to all kinds of things like external software packages, grid resource managers, data sources etc.
Triquetrum's initial contribution is based on Passerelle [2] which is in use since many years in several European synchrotrons that use workflows to automate experiment control, data acquisition and analysis.
An example
An interesting example is the fully automated MASSIF beamline of ESRF and EMBL at Grenoble, France. From the ESRF website [3]: "MASSIF-1 ... is a world leading unique facility for the fully automatic, high throughput characterisation and data collection from macromolecular crystals."
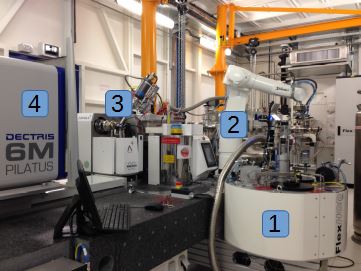

Crystallographic samples are stored at cryogenic temperatures in a Dewar sample container (1). A robotic arm (2) automatically picks each sample in sequence and transfers it to the measurement location (3) where X-ray diffraction patterns can be obtained by the detector (4). The sample must first be positioned and oriented very finely (micrometer precision) to be able to find the microscopic crystal and to capture the best possible diffraction images. The images must be processed by several dedicated programs to identify the molecular structure of e.g. proteins.
In the past such experiments and the data analysis had to be done as a sequence of manual actions, e.g. by visual inspection of the crystal position, passing image files to scripts and analysis programs etc. Now the whole sequence can be driven by one integrated workflow that automatically finds the optimal position and orientation of a sample, collects and analyses diffraction images and stores the results in the LIMS database.
Andrew McCarthy, Team Leader at EMBL, during a presentation of the Passerelle pilot:
"[executable] graphical workflow models definitely make it easier for us to collaborate on the automation of our MX experiments. After a limited introduction, they become almost self-documenting. The integration of Passerelle's execution traces with its graphical models allows our scientists to understand in detail what happened in each step of each experimental run."
Matthew Bowler, Staff Scientist at EMBL noted:
"The automatic routines developed are often able to locate crystals more effectively than the human eye and in many cases have obtained higher resolution data sets as all positions within a sample can be evaluated for diffraction quality.Initial experiments have been very successful, with users complimenting the accuracy, efficiency and simplicity of the facility."
Workflows?
The term workflow is used in many different contexts and with different meanings, e.g. ranging from a very abstract concept of "the way something is being done" to models and software to run processes in a deterministic way.
In the context of Triquetrum, a workflow is considered to be:
- a sequence of tasks to achieve a certain result
- that can be defined or recorded
- and can be executed or orchestrated in a deterministic way by a software tool
- potentially combining automated with interactive tasks
In enterprise environments, workflow systems are typically used to orchestrate tasks in business processes or document-oriented work, assigning activities to the right persons, doing follow-up on uncompleted tasks etc.
Scientific research is not about passing documents around or going through purchase approval processes. But software and computing resources have also become crucial in many scientific disciplines as the design, setup and execution of experiments is becoming more complex, the volume of obtained data that must be analysed is exploding and the demands for scientific "output productivity" are always increasing.
The integration of a workflow system in a platform for scientific software can bring many benefits:
- The steps in scientific processes are made explicitly visible in the workflow models (instead of being hidden inside program code or scripts). Such models can serve as a means to present, discuss and share scientific processes in communities with different skills-sets.
- Encapsulates technical services for automating complex processes in a scalable and maintainable way such as concurrent processing, support for integrating external programs and computing grids and their data transport and conversion needs, consistent error handling etc.
- Integrates execution tracing, provenance data, etc.
- Promotes modular solution design and reuse of model and software assets.
Some example applications:
- process control for scientific experiments: data acquisition, equipment control, integrated error recognition and recovery, monitoring & alarming
- in-silico experimentation through simulation or emulation
- (semi-)automated data reduction and analysis
- feedback between control & analysis in integrated workflows
- interactive assistance & support automation
Ptolemy II in Eclipse
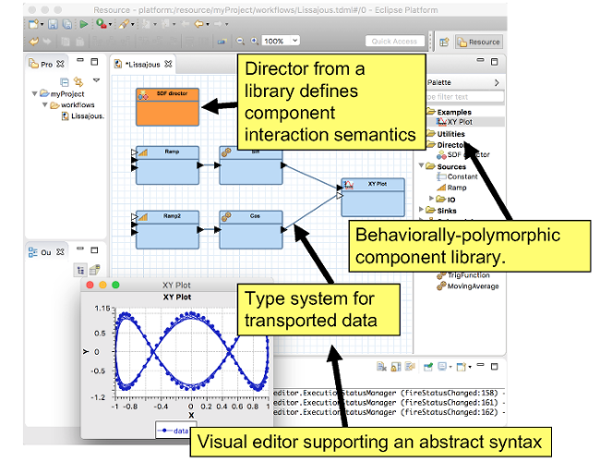
The core of Triquetrum is an integration of Ptolemy II [1] in an Eclipse and OSGi technology stack. Ptolemy II is an open source simulation and modeling tool from UC Berkeley. Originally intended for research and experimenting with system design techniques across engineering domains, it provides formalisms and tools for defining and executing hierarchical models of heterogeneous systems. Ptolemy II combines its actor-oriented architecture with a strong focus on concurrency, encapsulation and modularity. It comes with its own model design GUI called Vergil, based on Swing and a 2D graphical framework called Diva.
During its history of almost two decades it has been used internationally in varying domains, including in spin-offs for scientific workflow management like the Kepler Project [4] and Passerelle [2]. Triquetrum continues on the track started by Passerelle but now as an official Eclipse project linked to the Science IWG, and with a much closer integration and collaboration with the Ptolemy II team at UC Berkeley.
Triquetrum takes its name from the three cornered astronomical instrument held by Ptolemy. The name is meant to evoke the Model View Controller design pattern used in Ptolemy II.

Source: Wikipedia
Ptolemy II is an actor-oriented system where data moves between software components that are known as actors. Each actor encapsulates a well-defined function or responsibility, with a strict decoupling between actor implementations. Actors only communicate via ports. Actors execute concurrently and may be composed hierarchically.
The semantics of how the model is executed is known as the model of computation. Ptolemy II supports many different models of computation and these models of computation can be used at different levels of hierarchy.
The Ptolemy group at Berkeley intends to use Triquetrum to provide an Eclipse-based actor-oriented IoT development environment called Cape Code [5]. The key idea behind Cape Code is Accessors [6], which are actors that provide access to a service, sensor or actuator. Accessors are instantiated by an accessor host, which is a program. Accessor hosts use the accessor as if it were a local source or sink for data and commands. Currently, accessor hosts are implemented in Java, JavaScript in a web browser, Node.js, C and Rust, where a typical design flow is to develop an IoT system using Cape Code and then generate code for Node and then for C. Accessors themselves are written in small JavaScript files that define the ports, parameters and functions to be implemented on each accessor host. Accessor hosts then implement the functions used in the accessor definition. An accessor host is to IoT what a web browser is to the internet in that they both render remote services by locally executing a proxy for that service.
Status
Triquetrum has done its first release, as part of the Science 2016 release on October 21st. This release defines the core APIs with basic implementations and demonstrates the goals of the three lines of work of Triquetrum:
- The workflow runtime subsystem provides APIs and initial implementations for model repositories and workflow executions. The RCP editor is in a prototype stage, to demonstrate our ideas and the selected underlying frameworks, and to gather feedback from initial users.
- The task-based processing model provides a first iteration of the core APIs and some examples, e.g. of a basic integration in a workflow actor.
- For the third line of work, about adapters to integrate with external systems and other technical services, we demonstrate integration options to run C-Python scripts.
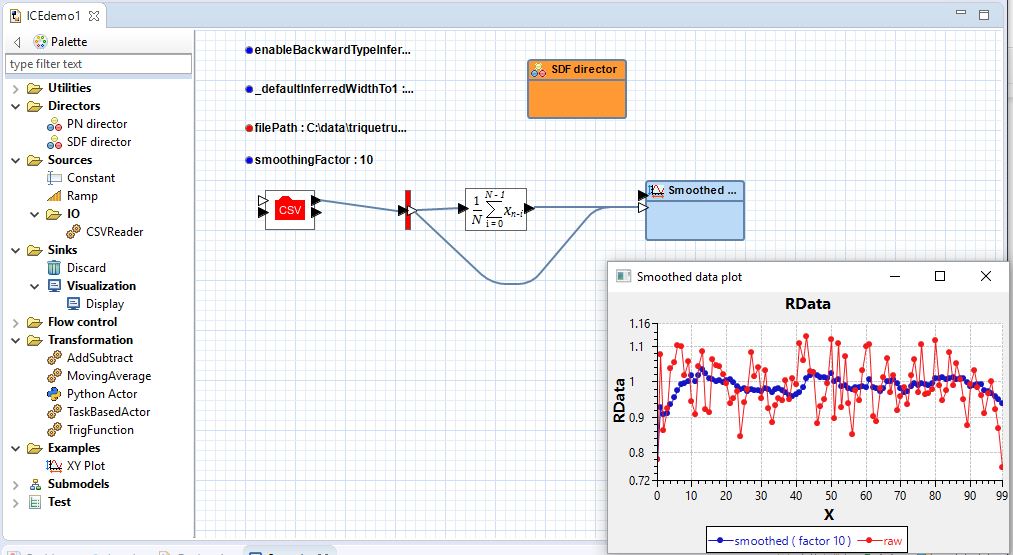
The screenshot shows some customized features of the Graphiti-based editor: a palette with a filtered treeview and custom actor icons based on SVG and Ptolemy II's custom icon designs.
Getting in touch
The Triquetrum wiki provides more information, e.g. how to get started with the RCP editor and an example of how to extend the system.
We use the triquetrum-dev mailing list, a low-volume blog and GitHub Issues to provide support and communicate with our community.
References
[1] http://ptolemy.eecs.berkeley.edu/ptolemyII/
[2] http://isencia.be/automation-in-a-scientific-research-area/
http://isencia.be/passerelle-edm-for-new-massif-beamlines/
https://github.com/eclipselabs/passerelle
[3] http://www.esrf.eu/home/news/general/content-news/general/massif-1-rise-of-the-robots-transforms-mx.html
[4] https://kepler-project.org/
[5] http://capecode.org
[6] http://accessors.org
About the Authors